Nine years ago, Craig Venter sequenced the first complete individual human genome - his own. Now, he's finally starting to decode what it means for his future.
In early 2006, Craig Venter received a worrying email concerning his genome. Amid the six billion letters-long sequence of the A, T, C and G base pairs that constitute the vocabulary of DNA, geneticists at his private research institute had discovered a single, errant "C". This mutation, called APOE 4, marked him for increased risk of cardiovascular disease, and a tripled likelihood of Alzheimer's. The higher cardiovascular risk came as little surprise for someone with a family history of early heart-attack deaths. But the Alzheimer's was unexpected. Advertisement
In one sense, few people could have been better prepared. In addition to having an entire institute's worth of geneticists on call, Venter is an undisputed pioneer in the field after leading his former company, Celera Genomics, in a fierce competition with the publicly funded Human Genome Project to produce the first complete human genome sequence. On June 26, 2000, in an uneasy truce that involved the intervention of then president Bill Clinton, the race was called as a tie.
In about a third of people who've come through [Human Longevity's clinic], we have found potentially lethal diseases
Read next
Yet in another respect he was quite fundamentally alone. Both Celera and the Human Genome Project used composite DNA from many different people - they revealed the framework of a human, but not the picture of any particular person. When Venter's entire genome was published in PLOS Biology on September 4, 2007, it stood in a contextual vacuum - the first, only, complete genome sequence of an individual in existence.
As a result, the role of even a recognised disease-related allele such as APOE 4 in the development of Alzheimer's was not, and still isn't, understood. Over half of those known to carry a single copy do not go on to develop the neurodegenerative disease. What other genetic variations might they have that protected them, and did Venter himself share them? What exactly did this APOE 4 variation mean for his future? Craig Venter had just become the first person in human history to stare the full extent of his own genetic determinism in the face. And he had next to no idea of what it all meant.
Sitting in a sunny office in the seaside town of La Jolla, California, surrounded by a collection of model ships and an old motorbike, Venter at 69 shows no sign of the deleterious effects predicted by his genomic code. In 2013, a brain scan cleared him of any build-up of the toxic beta amyloid plaques responsible for Alzheimer's. The sole symptom of the past eight years is perhaps a mellowing of a notoriously combative personality that exemplified the bitter rivalry of the human-genome race. The only medical concession Venter has made to his genomic risk factors has been to start taking statins, a commonly prescribed cholesterol-lowering drug. Hardly the revolution in predictive diagnostics and personalised treatment anticipated following the publication of the first complete individual genome. "Medically," Venter concludes of the $100m (£70m) sequencing effort, "it was not particularly useful."
On the computer screen in front of him, Venter shows WIRED how that's about to change. On the left is an ordinary photo of a middle-aged woman of mixed ethnicity, on the right a computer-generated likeness. With irises, skin and hair colour matched to the precise shade, it looks like the kind of photofit that police detectives - used to the greyscale grotesques produced by an imperfect human memory - could only dream of.
It was produced without seeing the subject's face.Read next

"From just the fingerprint on your pen, we can sequence your genome and identify how you look," Venter explains. "It's good enough to pick someone out of a ten-person line-up and it's getting better all the time." These prediction algorithms were developed at Venter's latest venture, biosciences startup Human Longevity, Inc (HLi) by measuring 30,000 datapoints from across the faces of a thousand volunteers, then using machine learning to identify patterns between their facial morphology and their entire genetic code. "We could take foetal cells from a mother's bloodstream, sequence the genome and give her a picture of what her future child will look like at 18," he says.
Eight years ago, Venter's genome couldn't even be used to tell you his eye colour. Yet modelling appearance is merely a visual demonstration, not the end goal, of HLi's big-data genomics. The aim is to predict your future. To explain why some people's cholesterol accumulates in their arteries to ultimately fatal levels, and others' does not. To identify which women are likely to develop breast or ovarian cancer in later life. And to understand why some people develop Alzheimer's and some continue to live a cognitively rich life at 90. "If you're a 55-year-old male you have a 30 per cent chance of dying before you reach 75," Venter says. "About a third of that is from cancer, and a third is from heart disease. If you're a female, your chance is about 20 per cent and two thirds of that is cancer. But these diseases are getting more and more predictable from your genome, and the theory is that if we can predict them, we can prevent them."
The trick is looking not just at single genes alone, but at complex patterns of interaction across the entirety of a person's DNA. "We're trying to wean the world off thinking of single-letter changes in a gene," says Venter. "There is no 'cancer gene'. A mutation in the BRCA1 or BRCA2 gene may increase your risk, but it cannot predict breast or ovarian cancer. But if you have family history as well, then your risk goes up to 90 per cent. These genes are being used as a surrogate for many other genetic factors."
Quite how many relevant genomic factors need to be considered has only recently been understood - the result of two linked discoveries that fundamentally changed our understanding of the human genome.Read next
In the early 2000s, the scientific picture was relatively simple. We know that every aspect of our bodies, from the colour of our hair and eyes to our natural muscle mass, is determined by the mix of hundreds of thousands of types of proteins from which our cells and the molecular machinery that sustains them are composed. Each type of protein, the hypothesis held, is produced according to instructions held in a single discrete section of DNA - a gene. Identification of genetic risk factors could therefore be straightforward. Know the gene, know the protein, know the problem. "Even brilliant geneticists wanted there to be hundreds of thousands of human genes so there would be one gene for everything," explains Venter. In 2004, an analysis of the Human Genome Project's results produced a revised estimate for the number of human genes from hundreds of thousands to fewer than 25,000 - fewer than a banana. The hypothesis went out the window.

Genomics pioneer Craig Venter in his office at Human Longevity, Inc in La Jolla, California
Then, in 2012, an international public research project, the Encyclopedia of DNA Elements (ENCODE), announced that they'd had a closer look at the remaining 98 per cent of our genome - the material outside known gene regions, which many had taken to dismissing as "junk". ENCODE claimed that as much as 80 per cent of our DNA actually encoded some function - including regulatory regions that control how the cell's molecular machinery reads a gene's protein-building instructions. The meaning of any one gene, how much and what type of protein it produces, may be determined by variations located millions of bases away. A single gene alone rarely identifies one protein, let alone the problem.
The significance of the 80 per cent figure is highly contested. The realisation that the gene-centric view neglects an essential part of the genome's story is not. Around 100 variations in non-protein-coding DNA have already been implicated in the development of various cancers. "From our first 10,000 genomes sequenced, we have found areas here where humans cannot have variation and still live," explains HLi's head of genomics, Amalio Telenti. "So these must code for something. We just don't know what it is yet."
What has been missing is the technology needed to process the enormous amount of information contained in a whole genome at scale. "The first complete genome cost $100 million and took nearly two years," says Venter. "That is not a replicable event. But a couple of hundred genomes aren't enough to learn anything. Sequencing accuracy, speed and cost had to change. Computing capabilities and storage costs had to change." In particular he points to how advances in machine learning allow for the extraction of complex patterns from a dataset far beyond human comprehension. "We've built the largest machine-learning team in genetics [led by former head of Google Translate, Franz Och]. Machine learning is essential because it looks at everything. It's not limited by hypotheses from the scientific literature or what gene-by-gene discovery has been made in the past." Read next
The final piece of the puzzle came in January 2014, with the release of the Illumina HiSeq X, the first high-throughput machine capable of sequencing an entire genome for as little as $1,000 - so long as you have $10 million up front and a rich enough supply of genetic material to keep the ten-unit system running full-time. In a laboratory beneath Venter's office, HLi has assembled the largest collection of HiSeq Xs in the world. Here, 24 fridge-sized machines - each named after a character from Star Wars- churn through 650-700 genomes a week, the equivalent of a genome sequence every 15 minutes. "On a weekly basis we generate about 60 terabytes of data just in this room," says head of genome sequencing William Biggs. "Facebook, on a monthly basis, uses 100 terabytes for all the photo uploads in the world. That kind of puts it in perspective."
Since being founded in 2013, HLi has used an initial $70 million in funding to sequence over 20,000 complete genomes, creating the largest whole-human-genome database in the world. By 2020, the aim is to reach one million. This genetic material comes from cancer centres across the US, from the UKTwin register of 12,000 twins, and from Alzheimer's patients in Brazil, alongside HLi's own trials and a $250 exome-sequencing service offered for clients of South African health-insurance provider Discovery Ltd. Although the exome includes just the two per cent of DNA in the protein-coding regions of our genes, this is still significantly more than other direct-to-consumer tests such as 23andMe, which detect only the presence of common alleles in known disease-related genes, such as BRCA1, BRCA2 and APOE. "The genetics community has been working under the notion that it's common variance that describes people," Venter says. "But it's not common alleles that determine your health and traits and physical outcomes, it's your rarer ones." Across the first 10,000 genomes analysed, HLi has discovered 82 million previously unseen types of genetic variation, adding to the most recent public database of 88 million, announced by the 1000 Genomes Project in October 2015. "We're expecting to have seen 500 million different types of variation by the time we've analysed 100,000 genomes," says Telenti. "Of the 170 million types discovered so far, 80 million have been seen only once, in a single person's genome."

24 HiSeq X machines sequence one genome every 15 minutes. Each unit is named after a Star Wars character
Not all this variation will be significant. Scientific opinion varies as to how much of our DNA actually enters into the body's programming, but all agree that much of it is redundant. Still, even non-coding variation has its uses. "We can tell identical twins apart by these slight variations," says Venter, which came in handy during one study of twin genomes when HLi found several people were entering their own DNA twice to claim double payment. "Now," he laughs, "we make sure you both turn up together."
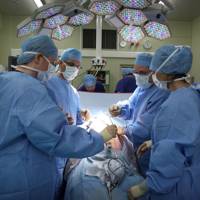
Every genetic sample sequenced by HLi is stored alongside its owner's phenotypic traits, from MRI scans showing the progression of neurodegeneration in the Brazilian Alzheimer's patients, to health questionnaires completed by Discovery's health-insurance clients. But by far the richest picture is being contributed by a select group of wealthy, health-conscious individuals, over 30 of whom so far have paid $25,000 to enter HLi's preventative-medicine centre, the Health Nucleus. "It's our own phenotyping clinic," says Venter. "We have MRI imaging of the brain and body which allows us to detect any tumour larger than two millimetres. We take 4D echocardiograms, to create a movie of your heart and measure every parameter in it. We also analyse 2,400 chemicals in the bloodstream to find correlations between these chemicals and the bacteria that create them."Read next
Age-related disease is responsible for two thirds of all deaths worldwide
Entrepreneurs, athletes, executives and celebrities are not paying $25,000 merely for the privilege of contributing to the lofty goals of scientific research. "In about a third of people who've come through, we have found potentially lethal diseases," says Venter. "We're criticised for doing these tests on healthy people, but clearly a third of them aren't, and they're pretty damn happy that we found what we did. In two people we discovered a greatly distended aorta - the first symptom would be a rupture and sudden death. One was a weightlifter with hypertension. That's not a good combination."
Walking through a fingerprint-scanner-secured door (genomic analysis being regrettably too slow for saliva-based security) into the Health Nucleus, WIRED is greeted by the fresh flowers, fruit and thick carpet that signal the transition to the customer-facing side of the business. Eight private rooms, one per client, surround the various diagnostics stations. There's the pink-lit MRI scanner, and the echocardiogram screen playing a looping movie of a patient's pulsating heart, but chief medical officer Brad Perkins becomes most enthusiastic when we enter a room containing what looks like nothing more than a patch of cheap linoleum. "This," he explains, "is our gait and balance test. It can actually provide an extraordinary amount of quantitative information about total neurological functioning. You can predict diseases like Parkinson's just from the gait."
On a television screen in one of the smartly furnished private rooms, consulting radiologist David Karow pulls up an image taken from one of the Health Nucleus's full-body MRI scans. It shows the cross section of a male's neck and shoulders, with a vibrant orange patch in the centre. Traditional greyscale MRI scans can be difficult to interpret - this one, clearly, is not. "Even a non-specialist could have picked that up," says Perkins. "That", is a stage one thymoma, a tumour growing beneath the breastbone, which can lead to a range of autoimmune diseases. "The patient had no symptoms other than a hoarse voice," says Perkins. "Fortunately, we caught it early and it was removed surgically, so he didn't even need radiotherapy. But he could have gone a couple of years with no more symptoms until all of a sudden it would have gone metastatic. At that point, it's no longer curable."
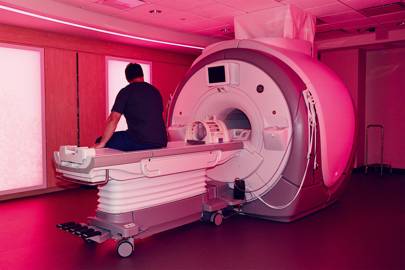
Although the MRI machine used at the Health Nucleus is standard - the same General Electric scanner used in hospitals all over the world - such clear, colourised images are not. "It used to be that the only way to see an image like this was to inject a dye into a patient's arteries," says Perkins. "About five per cent have a negative reaction and you really can't justify that risk in a screening situation." The Health Nucleus, he stresses, uses no dye in any scans. The images on the television are the result of a combination of new post-processing algorithms and tweaks to the MRI machine's imaging protocols. Karow, whose mother thrice had undetected tumours revealed by unrelated medical scans, is optimistic about such technology gaining a broader reach. "We're providing this at a high level right now, but we can use this data to come up with a more affordable programme for screening on a whole population level," he says. "As a radiologist it breaks my heart when you find a tumour and it's already metastatic. No one should die of metastatic cancer. It really is the case that cancer caught early is cancer cured." The pink-lit MRI scanner in the Health Nucleus. Pink is meant to relax anxious patientsRead next
For Venter it all comes back to the genome. All information collected in the Health Nucleus is anonymised and entered into the HLi master database alongside its owner's full genome sequence. "At the moment we're detecting these things from actual measurements, but we can take this data and use machine learning to look for new sites in the genome that would predict it," he says. "If we can find out what genetic variations are found in those patients with enlarged aortas, then next time we could predict it without even having to do the ultrasound."
Many later-life diseases will not be predictable from the genome. Lifestyle factors such as diet and exercise play an essential role in exacerbating, or reducing, the risk of cardiovascular disease and there are few genetic markers as predictive of early death as smoking. Just five to ten per cent of cancers are believed to be due to inherited factors, although according to HLi's head of cancer analysis, Barry Merryman, a surprisingly cheerful man given his specialism, our knowledge of these may change. "Currently we look at about 141 genes where there's strong evidence that a bad version greatly increases your chances of cancer, but we can't explain the majority of your inherited risk," he says. "We probably know only a third of it."
Still, a condition doesn't need to be inherited to be genetic. Ultimately, cancer always begins in the genome with the accumulation of mutations in the DNA of a single cell. "Part of what we're doing by sequencing people's genomes as well as their tumour, is finding new ways to predict from these changes that they're developing cancer in the first place," Merryman says. With many tens of trillions of cells in the body, the presence of such mutations won't be reflected in a standard DNA test, but these cancerous cells release fragments of their mutated DNA into the bloodstream, where the kind of high-throughput sequencing afforded by HLi's Illumina machines can potentially detect them. Illumina themselves recently formed a $100 million-funded spinoff company, Grail, to develop this non-invasive screening procedure.
Merryman has further reason to be optimistic. Beyond early detection, whole-genome analysis is also proving to be a powerful cancer-fighting tool. By comparing the full genome of both tumour and patient, HLi are able to identify exactly how mutations have altered the proteins covering the outside of cancerous cells. Replicating this specific protein structure and delivering it to the patient can train their immune system to recognise it as alien and to attack the tumour, just as inert viruses are used to train the immune system to fight off a live infection more quickly. "We had a patient with neck cancer, where the tumour had gone metastatic and wasn't responding to any treatment," says Merryman. "So we developed a vaccine from the mutated proteins in her tumour, and boosted all her immune cells. After a month we've already seen an 80 per cent reduction in total volume."Read next
Standing in the East Room of the White House in 2000, Venter announced that the sequencing of the first full human genome would bring about "the potential to reduce the number of cancer deaths to zero in our lifetimes." Just 100 hundred years prior, cancer, combined with heart disease, stroke, Alzheimer's and other age-related conditions that HLi aims to predict and prevent, were collectively responsible for less than a third of deaths. The majority of the population died of external attack by infectious disease long before their bodies' own self-destructive tendencies kicked in. Now, age-related disease is responsible for two thirds of all deaths worldwide, around 100,000 deaths a day. "In the last century, life expectancy has risen by about 30 years thanks to antibiotics and improved hygiene," explains University of Illinois professor and specialist in the demography of ageing, Jay Olshansky, "But there was a price to pay for that extended life - a rise in heart disease, cancer, stroke and Alzheimer's, all of which are the killers today."
These conditions place an enormous burden on the healthcare systems and economies of developed nations. The National Health Service's spend per head is now over four and a half times higher for the over-65s than for the under-25s. By 2050, over-65s are forecast to comprise more than 21 per cent of the global population, up from 12 per cent now. Alzheimer's and other forms of dementia loom large. In 2015, due to the chronic nature of the disease, the cost of care in the UK was over £26 billion, two thirds of which came from the families and friends of patients. Worldwide, the number of dementia sufferers is expected to double by 2036.
With our first 20,000 genomes we have just completed the first lap in what will be a very long race
Extending longevity one condition at a time is a flawed approach, Olshansky argues. "If you cured cancer, you only gain about three and a half years of life expectancy. It takes seven years for the risk of everything that is unpleasant about ageing to double. If we could delay ageing itself by just seven years, then your risk of heart disease, cancer, stroke and Alzheimer's would be reduced by 50 per cent throughout the remainder of your life."
Ageing comes next, explains HLi co-founder and president of the cellular-therapeutics division, Robert Hariri. "To have an impact on longevity we must first eradicate the causes of premature death, and the biggest factor here is cancer. Next is to identify and address the causes of degeneration in ageing." For this, HLi's focus is on stem cells, the undifferentiated cells deployed by the body to repair cellular damage. "At 60, you have a tiny fraction of the stem cells in your bone marrow that you had during your youth," says Hariri. "If we can keep recharging an individual's supply as they age we should be able to restore some youthful functionality and disease resistance. We're going to be looking to bank a baby's stem cells at birth, identify genetic issues, edit that genome to correct the abnormality and then give those cells back to them."
The topic of human genetic modification has recently been injected with particular urgency. As WIRED visits HLi in early December, scientists are gathering in Washington, DC, to discuss a new genetic-modification technique, CRISPR/Cas9. In April 2015, geneticists at Sun Yat-sen University in China announced they had used it to perform the first direct genetic edits to human embryos.Advertisement
The embryos were non-viable and the edits largely unsuccessful, yet the possibilities sent shockwaves through the scientific world. Unlike the re-implantation of a person's genetically modified adult cells - already used in some experimental therapies - such direct embryonic modifications proliferate throughout the developing body, making them inheritable by future offspring. The obvious question, as HLi begins to unravel the meaning of our genetic code, is when they plan to start editing it. Could we cut nasty, harmful predispositions out of the human species altogether?
Venter pulls hard on the brakes. "There are diseases where it would clearly be beneficial, but there's much more concern about making inheritable changes, because genes don't have just one function," he says. "You think you're fixing that one disease, and it could be causing others, in you or in your offspring." HLi's understanding may be advancing fast, but the kind of knowledge necessary for extensive modification of the genome is still a distant prospect. "We are making discoveries every day that surprise us because nobody has had these data sets before," he says, "Nobody has been able to ask these questions before. With our first 20,000 genomes we have just completed the first lap in what will be a very long race."
Kathryn Nave is a contributing editor to WIRED. She wrote about Shenzhen's tech markets in 04.16