404 Not Found

Every shrub, every tree—
if one has not forgotten
where they were planted—
has beneath the fallen snow
some vestige of its form.
Try to remember what you were looking for...

A thousand ages in thy sight
Are like an evening gone,
Short as the watch that ends the night
Before the rising sun.

…The Sea of Faith
Was once, too, at the full, and round earth’s shore
Lay like the folds of a bright girdle furled.
But now I only hear
Its melancholy, long, withdrawing roar,
Retreating, to the breath
Of the night-wind, down the vast edges drear
And naked shingles of the world.Ah, love, let us be true
To one another! for the world, which seems
To lie before us like a land of dreams,
So various, so beautiful, so new,
Hath really neither joy, nor love, nor light,
Nor certitude, nor peace, nor help for pain;
And we are here as on a darkling plain
Swept with confused alarms of struggle and flight,
Where ignorant armies clash by night.Matthew Arnold, “Dover Beach”

Action is transitory—a step, a blow,
The motion of a muscle—this way or that—’Tis done, and in the after vacancy
We wonder at ourselves like men betrayed:Suffering is permanent, obscure and dark,
And shares the nature of infinity.

Come, fill the Cup, and in the fire of Spring
The Winter Garment of Repentance fling:
The Bird of Time has but a little way
To fly—and lo! the Bird is on the Wing.

The Worldly Hope men set their Hearts upon
Turns Ashes—or it prospers; and anon,
Like Snow upon the Desert’s dusty Face,
Lighting a little hour or two—is gone.

Alike for those who for To-day prepare,
And those that after some To-morrow stare,
A Muezzín from the Tower of Darkness cries
“Fools! your Reward is neither Here nor There.”

Why, all the Saints & Sages who discuss’d
Of the Two Worlds so wisely—they are thrust
Like foolish Prophets forth; their Words to Scorn
Are scatter’d, & their Mouths are stopt with Dust.
Oh, come with old Khayyam, and leave the Wise
To talk; one thing is certain, that Life flies;
One thing is certain, and the Rest is Lies;
The Flower that once has blown for ever dies.

Myself when young did eagerly frequent
Doctor and Saint, and heard great Argument
About it and about: but evermore
Came out by the same Door as in I went.

With them the seed of Wisdom did I sow,
And with mine own hand wrought to make it grow;
And this was all the Harvest that I reap’d—
“I came like Water, and like Wind I go.”

Up from Earth’s Center through the Seventh Gate
I rose, and on the Throne of Saturn sate;
And many Knots unravel’d by the Road;
But not the Knot of Human Death and Fate.

There was a Door to which I found no Key;
There was a Veil past which I could not see:
Some little talk awhile of Me and Thee
There seemed—and then no more of Thee & Me.

Ah, fill the Cup:—what boots it to repeat
How Time is slipping underneath our Feet:
Unborn To-morrow, and dead Yesterday,
Why fret about them if To-day be sweet!

The Ball no Question makes of Ayes and Noes,
But Right or Left as strikes the Player goes;
And He that toss’d Thee down into the Field,
He knows about it all—He knows—He knows!

The Moving Finger writes; &, having writ,
Moves on: nor all thy Piety nor Wit
Shall lure it back to cancel half a Line,
Nor all thy Tears wash out a Word of it.

Oh Thou, who didst with Pitfall & with Gin
Beset the Road I was to wander in,
Thou wilt not with Predestination round
Enmesh me, and impute my Fall to Sin?

None answer’d this; but after Silence spake
A Vessel of a more ungainly Make:
“They sneer at me for leaning all awry;
What? did the Hand then of the Potter shake?”

Ah, Moon of my Delight who know’st no wane,
The Moon of Heav’n is rising once again:
How oft hereafter rising shall she look
Through this same Garden after me—in vain!

Ah, Love! could thou and I with Fate conspire
To grasp this sorry Scheme of Things entire
Would not we shatter it to bits—and then
Re-mould it nearer to the Heart’s Desire!

Et in Arcadia ego

Decay is inherent in all compound things.
Work out your own salvation with diligence.Last words of the Buddha

Uh oh!
—Gwern (first words)

What do you mean,
404 Not Found‽

I’m as confused as you are.

The Great Work goes on.
Do not summon up that which you cannot put down.





GIGO.














With every new spring
the blossoms speak not a word
yet expound the Law—
knowing what is at its heart
by the scattering storm winds.
Anything you post on the internet will be there as long as it’s embarrassing and gone as soon as it would be useful.
With the people of the past
How I wish I could share the beauty
Of these cherry blossoms.
Have each of us across the years
Left behind this very thought?
For Books are not absolutely dead things, but doe contain a potencie of life in them to be as active as that soule was whose progeny they are; nay they do preserve as in a violl the purest efficacie and extraction of that living intellect that bred them. I know they are as lively, and as vigorously productive, as those fabulous Dragons teeth; and being sown up and down, may chance to spring up armed men. And yet on the other hand, unlesse warinesse be us’d, as good almost kill a Man as kill a good Book; who kills a Man kills a reasonable creature, Gods Image; but hee who destroyes a good Booke, kills reason it selfe, kills the Image of God, as it were in the eye. Many a man lives a burden to the Earth; but a good Booke is the pretious life-blood of a master spirit, imbalm’d and treasur’d up on purpose to a life beyond life. ‘Tis true, no age can restore a life, whereof perhaps there is no great losse; and revolutions of ages do not oft recover the losse of a rejected truth, for the want of which whole Nations fare the worse. We should be wary therefore what persecution we raise against the living labours of publick men, how we spill that season’d life of man preserv’d and stor’d up in Books; since we see a kinde of homicide may be thus committed, sometimes a martyrdome, and if it extend to the whole impression, a kinde of massacre, whereof the execution ends not in the slaying of an elementall life, but strikes at that ethereall and fift essence, the breath of reason it selfe, slaies an immortality rather then a life.
Milton
“The horror of that moment”, the King went on, “I shall never never forget!” “You will, though”, the Queen said, “if you don’t make a memorandum of it.”
Alice in Wonderland, Lewis Carroll
The great globe reels in the solar fire,
Spinning the trivial and unique away.
(How all things flash! How all things flare!)
...Time is the school in which we learn
Time is the fire in which we burn...
(This is the school in which we learn ...)
What is the self amid this blaze?
What am I now that I was then
Which I shall suffer and act again,
The theodicy I wrote in my high school days
Restored all life from infancy,
The children shouting are bright as they run
(This is the school in which they learn ...)
Ravished entirely in their passing play!
(...that time is the fire in which they burn.)Delmore Schwartz (“Calmly We Walk through This April’s Day”)
I was decimated. To program any more would be pointless. My programs would never live as long as The Trial. A computer will never live as long as The Trial. ...What if Amerika was only written for 32-bit PowerPC?
_why
Who alive can say,
“Thou art no Poet—may’st not tell thy dreams”?
Since every man whose soul is not a clod
Hath visions, and would speak, if he had loved,
And been well nurtured in his mother tongue.John Keats (The Fall of Hyperion: A Dream I 11-5)
Expressions come to an end somewhere.
Wittgenstein (Philosophical Investigations)
I cannot tell how the truth may be;
I say the tale as ’twas said to me.Sir Walter Scott
What has been done, thought, written, or spoken is not culture; culture is only that fraction which is remembered.
Gary Taylor (The Clock of the Long Now)
What is the door for, opening or closing? What do you think? Don’t look at me like that. This is a very important question for me. Especially in here, the deepest underground, there are so many doors.
Well, well, well. What do you think?
If you’d like to say, “I don’t know”, how about this instead? “The door should be opened by force.” That’s the reason. That’s why I’m here.Ryoji Kaji (2015: The Last Year of Ryoji Kaji)
What is wanted is not the will to believe, but the wish to find out, which is the exact opposite.
Bertrand Russell (“Free Thought and Official Propaganda”)
There is no point in using the word ‘impossible’ to describe something that has clearly happened.
Douglas Adams (Dirk Gently’s Holistic Detective Agency)
There is no view from nowhere.
Thomas Nagel
There is nothing so useless as doing efficiently that which should not be done at all.
Peter Drucker
There must have been a moment at the beginning, where we could have said no.
Somehow we missed it.
Well, we’ll know better next time.Guildenstern (Rosencrantz & Guildenstern Are Dead, Tom Stoppard)
Therefore, in emptiness, no form,
No feelings, perceptions, impulses, consciousness;
No eyes, no ears, no nose, no tongue, no body, no mind;
No color, no sound, no smell, no taste, no touch, no object of mind...Heart Sutra
What you are now is the result of what you were.
What you will be tomorrow will be the result of what you are now.
The consequences of an evil mind will follow you like the cart follows the ox that pulls it.
The consequences of a purified mind will follow you like your own shadow.
No one can do more for you than your own
purified mind—no parent, no relative, no friend, no one.
A well-disciplined mind brings happiness.the Dhammapada
To tell Diomedes’ story he [Homer] doesn’t think
He has to start with the death of the hero’s uncle,
Or start, in telling about the Trojan War,
By telling us how Helen came out of an egg.
He goes right to the point and carries the reader
Into the midst of things, as if known already;
And if there’s material that he despairs of presenting
So as to shine for us, he leaves it out;
& he makes his whole poem one. What’s true, what’s invented,
Beginning, middle, and end, all fit together.Horace (Ars Poetica)
Perfection is our goal. Excellence will be tolerated.
J. Yahl
Philosophy in any case always comes on the scene too late...When philosophy paints its gloomy picture then a form of life has grown old. It cannot be rejuvenated by the gloomy picture, but only understood.
Only when the dusk starts to fall does the owl of Minerva spread its wings and fly.
G. W. F. Hegel
I went down into Shropshire to look at that famous library [of the late Lord Acton] before it was removed to Cambridge. There were shelves on shelves of books on every conceivable subject—Renaissance Sorcery, the Fueros of Aragon, Scholastic Philosophy, the Growth of the French Navy, American Exploration, Church Councils. The owner had read them all, and many of them were full in their margins with cross-references in pencil. There pigeon-holed desks and cabinets with literally thousands of compartments, into each of which were sorted little white slips with references to some particular topic, so drawn up (so far as I could see) that no one but the compiler could easily make out the drift of the section. I turned over one or two from curiosity—one was on early instances of a sympathetic feeling for animals, from Ulysses’ old dog in Homer downward. Another seemed to be devoted to a collection of hard words about stepmothers in all national literatures, a third seemed to be about tribal totems. Arranged in the center of the room was a sort of block or altar composed entirely of unopened parcels of new books from continental publishers. All had arrived since Lord Acton’s health began to break up. These volumes were apparently coming in at the rate of ten or so per week, and the purchaser had evidently intended to keep pace with the accumulation, to read them all, and to work their results into his vast thesis—whatever it was. For years, apparently, he had been endeavoring to keep up with everything that was being written—a Sisyphean task. Over all there were brown Holland sheets, a thin coating of dust, the moths dancing in the pale September sun. There was a faint aroma of mustiness, proceeding from thousands of 17th & 18th-century books in a room that had been locked up since the owner’s death. I never saw a sight that more impressed on me the vanity of human life and learning.
Sir Charles Oman (1939)
The road to wisdom?—Well, it’s plain
and simple to express:
Err
and err
and err again
but less
and less
and less.Piet Hein
The single biggest problem in communication is the illusion that it has taken place.
George Bernard Shaw
The sky above the port was the color of television, tuned to a dead channel.
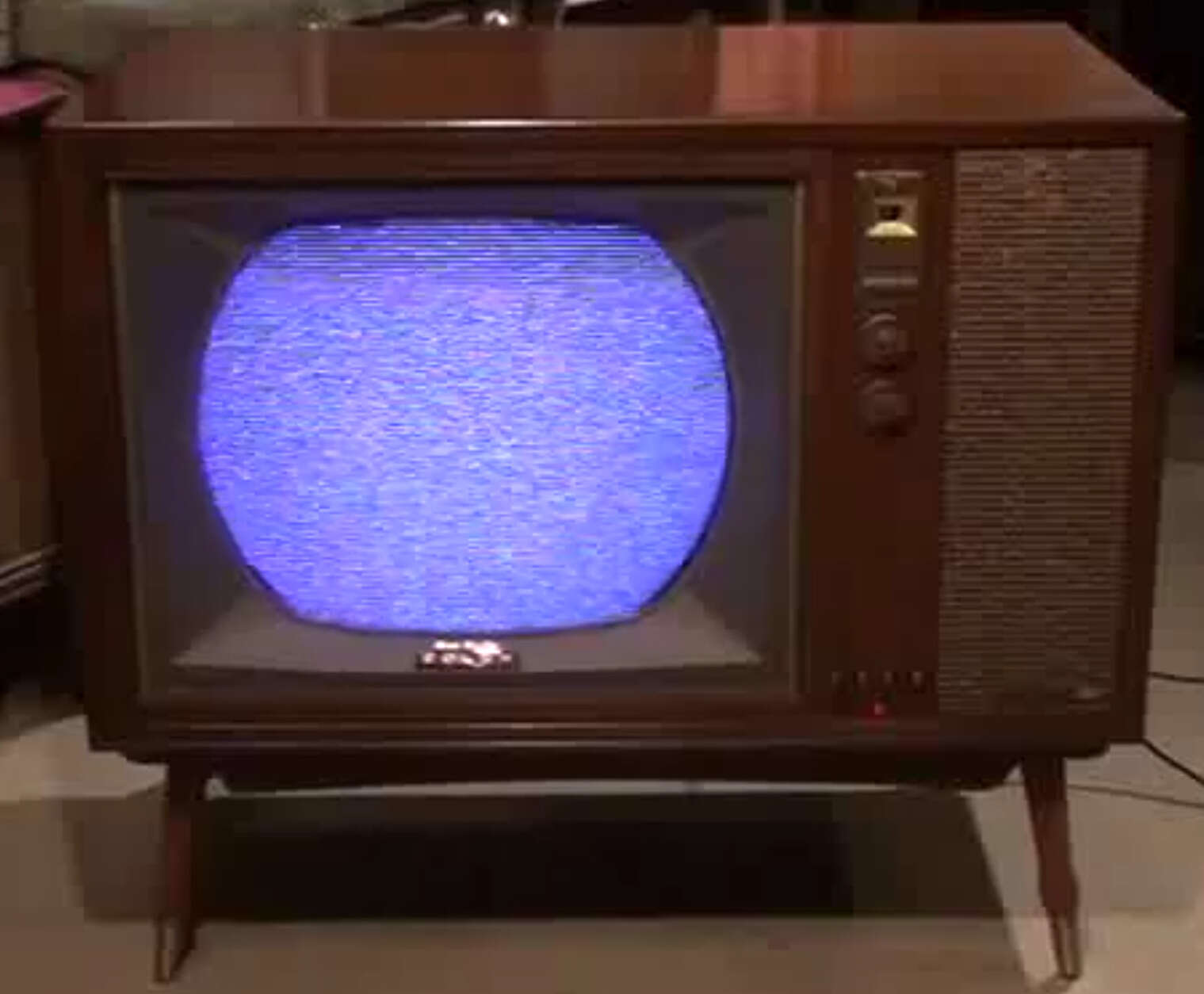
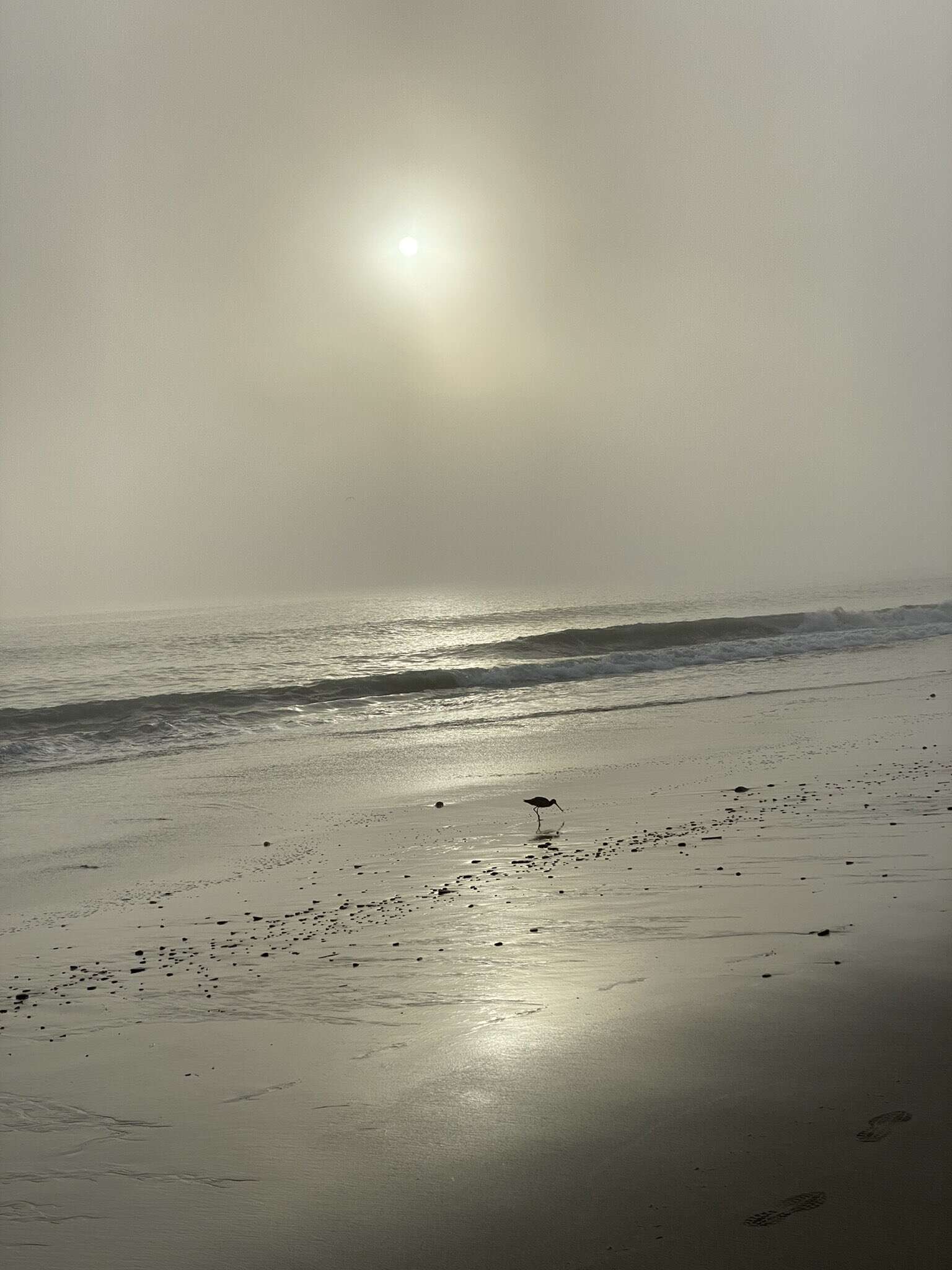



The software entity is constantly subject to pressures for change. Of course, so are buildings, cars, computers. But manufactured things are infrequently changed after manufacture; they are superseded by later models, or essential changes are incorporated into copies of the same basic design. Callbacks of automobiles are really quite infrequent; field changes of computers somewhat less so. Both are much less frequent than modifications to fielded software.
In part, this is so because the software of a system embodies its function, and the function is the part that most feels the pressures of change. In part it is because software can be changed more easily—it is pure thought-stuff, infinitely malleable.
Fred Brooks (The Mythical Man-Month)
The soul has no assignments, neither cooks
Nor referees: it wastes its time. It wastes its time.
Here in this enclave there are centuries
For you to waste: the short and narrow stream
Of life meanders into a thousand valleys
Of all that was, or might have been, or is to be.
The books, just leafed through, whisper endlessly.Randall Jarrell (“A Girl in a Library”)
We think that powerful and lifeful movement is impossible without differences—“true conformity” is possible only in the cemetery.
Joseph Stalin (1912)
We try things. Occasionally they even work.
Parson Gotti (Erfworld, Rob Balder)
We wanted the best, but it turned out like always.
Viktor Chernomyrdin
We will encourage you to develop the 3 great virtues of a programmer: laziness, impatience & hubris.
Larry Wall & Randal Schwartz (Programming Perl)
What do they think has happened, the old fools
To make them like this? Do they somehow suppose
It’s more grown-up when your mouth hangs open and drools
And you keep on pissing yourself, and can’t remember
Who called this morning? Or that, if they only chose
They could alter things back to when they danced all night
Or went to their wedding, or sloped arms some September?
Or do they fancy there’s really been no change
And they’ve always behaved as if they were crippled or tight
Or sat through days of thin continuous dreaming
Watching light move? If they don’t (and they can’t), it’s strange
Why aren’t they screaming?Philip Larkin (“The Old Fools”)
What is man, that thou art mindful of him? and the son of man, that thou visitest him?
Psalm 8:4
Whatever became of the moment when one first knew about death? There must have been one. A moment. In childhood. When it first occurred to you that you don’t go on forever. Must have been shattering, stamped into one’s memory. And yet, I can’t remember it. It never occurred to me at all.
Rosencrantz & Guildenstern Are Dead (Tom Stoppard)
Whatsoever thy hand findeth to do, do it with thy might; for there is no work, nor device, nor knowledge, nor wisdom, in the grave, whither thou goest.
Qoheleth
When I look upon
the rich sheen of summer hair
in my new writing brush,
I am saddened: by a deer,
drawn at night to a hunter’s torch.Shōtetsu (‘Summer Writing Brush’)
When I pronounce the word Future,
the first syllable already belongs to the past.When I pronounce the word Silence,
I destroy it.When I pronounce the word Nothing,
I make something no nonbeing can hold.Wisława Szymborska (“The Three Oddest Words”)
When a creature has developed into one thing, he will choose death rather than change into his opposite.
Scytale (Dune Messiah, Frank Herbert)
When a thing is placed
A shadow of autumn
Appears there.Kyoshi Takahama
When autumn winds blow
not one leaf remains
the way it was.Togyu
When copies are super abundant, copies are worthless...When copies are super abundant, stuff which can’t be copied becomes scarce and valuable.
...Findability—Whereas the previous generative qualities reside within creative digital works, findability is an asset that occurs at a higher level in the aggregate of many works. A zero price does not help direct attention to a work, and in fact may sometimes hinder it. But no matter what its price, a work has no value unless it is seen; unfound masterpieces are worthless. When there are millions of books, millions of songs, millions of films, millions of applications, millions of everything requesting our attention—and most of it free—being found is valuable.
Kevin Kelly (“Better than Free”)
When human beings found out about death
They sent the dog to Chukwu with a message:
They wanted to be let back to the house of life.
They didn’t want to end up lost forever
Like burnt wood disappearing into smoke
And ashes that get blown away to nothing.
Instead, they saw their souls in a flock at twilight
Cawing and headed back for the same old roosts
(The dog was meant to tell all this to Chukwu)...Seamus Heaney (“A Dog Was Crying Tonight in Wicklow Also”)
When it blows,
The mountain wind is boisterous,
But when it blows not,
It simply blows not.Ikkyu
When setting out upon your way to Ithaca,
wish always that your course be long,
full of adventure, full of lore.Constantine P. Cavafy (“Ithaka”)
When the assembly could not reply and if I had been Nan-ch’üan, I would have released the cat, since the assemblage had already said they could not answer.
An old Master has said: “In expressing full function, there are no fixed methods.”Dōgen (Shōbōgenzō)
When you have eliminated the impossible, whatever remains is often more improbable than your having made a mistake in one of your impossibility proofs.
Steven Kaas
While I gazed out,
barely conscious that I too
was growing old,
how many times have blossoms
scattered on the spring wind?Fujiwara no Teika
While life is yours, live joyously;
None can escape Death’s searching eye:
When once this frame of ours they burn,
How shall it e’er again return?Carvaka
While you live, shine.
Don’t suffer anything at all;
Life exists only a short while
And time demands its toll.Seikilos epitaph
Who in the future
Will recall me in the scent
Of orange blossoms
When I, too, shall have become
A person of long ago?Fujiwara no Shunzei
Whoever does not know how to hit the nail on the head should be entreated to not hit the nail at all.
Friedrich Nietzsche
No place affords a more striking conviction of the vanity of human hopes than a public library.
Dr. Samuel Johnson
No shelter in sight
to give my pony a rest
and brush off my sleeves—
in the fields around Sano Ford
on a snowy evening.Fujiwara no Teika
On a journey, ill—
my dream goes wandering
over withered fields.Bashō
On autumn nights
The dew feels particularly
Cold,
When in every patch of grass
The insects sorrow.Anonymous
On the one hand, information wants to be expensive because it’s so valuable. The right information in the right place just changes your life. On the other hand, information wants to be free, because the cost of getting it out is lower and lower all the time. So you have these 2 things fighting against each other.
Stewart Brand to Steve Wozniak (1984)
On two occasions I have been asked,—‘Pray, Mr. Babbage, if you put into the machine wrong figures, will the right answers come out?’ In one case a member of the Upper, and in the other a member of the Lower, House put this question. I am not able rightly to apprehend the kind of confusion of ideas that could provoke such a question.
Charles Babbage (Passages from the Life of a Philosopher)
The river darkens on an autumn night
And the waves subside as if to sleep.
I drop a line into the water
But the sleepy fish won’t bite.The empty boat and I return
Filled with our catch of moonlight.Yi Jung
C’est pire qu’un crime; c’est une faute.
[“It’s worse than a crime; it’s a mistake.”]Talleyrand (on the killing of the Duc D’Enghien)
“Are you coming to bed?”
“I can’t. This is important.”
“What?”
“Someone is wrong on the Internet.”XKCD (“Duty Calls”)
“Think of the past!”—
so the moonlight seems to say,
itself a remnant
of autumns long since gone,
that I could never know.Fujiwara no Teika
‘We name things & then we can talk about them: can refer to them in talk.’—As if what we did next were given with the mere act of naming. As if there were only one thing called ‘talking about a thing’. Whereas in fact we do the most various things with our sentences. Think of exclamations alone, with their completely different functions.
Water!
Away!
Ow!
Help!
Fine!
No!
Are you still inclined to call these words ‘names of objects’?...Wittgenstein (Philosophical Investigations)
“You see this goblet?” asks Achaan Chaa, the Thai meditation master. “For me this glass is already broken. I enjoy it; I drink out of it. It holds my water admirably, sometimes even reflecting the sun in beautiful patterns. If I should tap it, it has a lovely ring to it. But when I put this glass on the shelf and the wind knocks it over or my elbow brushes it off the table and it falls to the ground and shatters, I say, ‘Of course’.
When I understand that the glass is already broken, every moment with it is precious.”
Mark Epstein (Thoughts Without a Thinker)
What makes the desert beautiful is that it hides, somewhere, a well.
Antoine de Saint Exupéry (Le Petit Prince)
Does not a white bird
Feel within her heart forlorn?
The blue of the sky
The blue of the sea. Neither
Stains her, between them she floats.Wakayama Bokusui
Earth and metal...
although my breathing ceases
time and tide go on.Atsujin
For a successful technology, reality must take precedence over public relations, for nature cannot be fooled.
Richard Feynman (“Appendix F: Personal Observations on the Reliability of the Shuttle”)
For we can always see and feel much that the people in old photos and newsreels could not:
that their clothing and automobiles were old-fashioned,
that their landscape lacked skyscrapers and other contemporary buildings,
that their world was black
and white
and haunting
and gone.Kramer et al


For ‘Tragedy’ [τραγωδία] and ‘Comedy’ [τρυγωδία] come to be out of the same letters.
Democritus
From long ago
I have been traveling,
yet never arriving.
Even the old have far to go:
for that is the way with this Way.Shōtetsu (“Reminiscing”)

From too much love of living,
From hope and fear set free,
We thank with brief thanksgiving
Whatever gods may be
That no man lives for ever;
That dead men rise up never;
That even the weariest river
Winds somewhere safe to sea.Then star nor sun shall waken,
Nor any change of light;
Nor sound of waters shaken,
Nor any sound or sight;
Nor wintry nor vernal,
Nor days, nor things diurnal;
Only the sleep eternal
In an eternal night.Algernon C. Swinburne (“The Garden of Proserpine”)
Do not seek to follow in the footsteps of the old masters.
Seek what they sought.Bashō
We cross our bridges when we come to them, and burn them behind us—with nothing to show for our progress except a memory of the smell of smoke, and a presumption that once our eyes watered.
Guildenstern (Rosencrantz & Guildenstern Are Dead, Tom Stoppard)
I returned, and saw under the sun, that the race [is] not to the swift, nor the battle to the strong, neither yet bread to the wise, nor yet riches to men of understanding, nor yet favour to men of skill; but time and chance happeneth to them all.
Qoheleth
One does not care to acknowledge the mistakes of one’s youth.
Char Aznable
But where shall wisdom be found?
And where is the place of understanding?
Man knoweth not the price thereof;
neither is it found in the land of the living
...for the price of wisdom is above rubies.Book of Job, 28:12–28
Sampling can show the presence of knowledge but not the absence.
Show me the person who doesn’t die—
death remains impartial.
I recall a towering man
who is now a pile of dust.
The World Below knows no dawn
though plants enjoy another spring;
those visiting this sorrowful place
the pine wind slays with grief.Han-Shan
Empty-handed I entered the world
Barefoot I leave it.
My coming, my going—
Two simple happenings
That got entangled.Kozan Ichikyo
Evening cherry-blossoms:
I slip inkstone back into kimono
this one last time.Kaisho
Every drop of blood has great talent; the original cellule seems identical in all animals, and only varied in its growth by the varying circumstance which opens now this kind of cell and now that, causing in the remote effect now horns, now wings, now scales, now hair; and the same numerical atom, it would seem, was equally ready to be a particle of the eye or brain of man, or of the claw of a tiger...The man truly conversant with life knows, against all appearances, that there is a remedy for every wrong, and that every wall is a gate.
Ralph Waldo Emerson
Einstein argued that there must be simplified explanations of nature, because God is not capricious or arbitrary.
No such faith comforts the software engineer.Fred Brooks
Every year without knowing it I have passed the day
When the last fires will wave to me
And the silence will set out
Tireless traveler
Like the beam of a lightless starThen I will no longer
Find myself in life as in a strange garment
Surprised at the earth
And the love of one woman
And the shamelessness of men
As today writing after three days of rain
Hearing the wren sing and the falling cease
And bowing not knowing to what.W. S. Merwin (“For the Anniversary of My Death”)

There is a line of Verlaine I shall not recall again,
There is a nearby street forbidden to my step,
There is a mirror that has seen me for the last time,
There is a door I have shut until the end of the world.
Among the books in my library (I have them before me)
There are some I shall never reopen.
This summer I complete my 50th year:
Death reduces me incessantly.Jorge Luis Borges (“Limits”)
Everybody has got to die, but I always believed an exception would be made in my case.
William Saroyan (letter written to his survivors)
What Song the Syrens sang, or what name Achilles assumed when he hid himself among women, though puzling Questions are not beyond all conjecture. What time the persons of these Ossuaries entred the famous Nations of the dead, and slept with Princes and Counsellours, might admit a wide resolution. But who were the proprietaries of these bones, or what bodies these ashes made up, were a question above Antiquarism. Not to be resolved by man, nor easily perhaps by spirits, except we consult the Provinciall Guardians, or tutellary Observators. Had they made as good provision for their names, as they have done for their Reliques, they had not so grosly erred in the art of perpetuation. But to subsist in bones, and be but Pyramidally extant, is a fallacy in duration. Vain ashes, which in the oblivion of names, persons, times, and sexes, have found unto themselves, a fruitlesse continuation, and only arise unto late posterity, as Emblemes of mortall vanities; Antidotes against pride, vain-glory, and madding vices. Pagan vain-glories which thought the world might last for ever, had encouragement for ambition, and finding no Atropos unto the immortality of their Names, were never dampt with the necessity of oblivion. Even old ambitions had the advantage of ours, in the attempts of their vain-glories, who acting early, and before the probable Meridian of time, have by this time found great accomplishment of their designes, whereby the ancient Heroes have already out-lasted their Monuments, and Mechanicall preservations. But in this latter Scene of time we cannot expect such Mummies unto our memories, when ambition may fear the Prophecy of Elias, and Charles the Fifth can never hope to live within two Methusela’s of Hector.
Sir Thomas Browne (Hydriotaphia, Urn Burial)
Nature hath furnished one part of the Earth, and man another. The treasures of time lie high, in Urnes, Coynes, and Monuments, scarce below the roots of some vegetables. Time hath endlesse rarities, and shows of all varieties; which reveals old things in heaven, makes new discoveries in earth, and even earth it self a discovery. That great Antiquity America lay buried for a thousand years; and a large part of the earth is still in the Urne unto us.
Sir Thomas Browne (Hydriotaphia, Urn Burial)
But the iniquity of oblivion blindely scattereth her poppy, and deals with the memory of men without distinction to merit of perpetuity. Who can but pity the founder of the Pyramids? Herostratus lives that burnt the Temple of Diana, he is almost lost that built it; Time hath spared the Epitaph of Adrians horse, confounded that of himself. In vain we compute our felicities by the advantage of our good names, since bad have equall durations; and Thersites is like to live as long as Agamenon, [without the favour of the everlasting Register:] Who knows whether the best of men be known? or whether there be not more remarkable persons forgot, then any that stand remembred in the known account of time? without the favour of the everlasting Register the first man had been as unknown as the last, and Methuselah’s long life had been his only Chronicle.
Sir Thomas Browne (Hydriotaphia, Urn Burial)
The strong do what they will, and the weak suffer what they must.
Thucydides (“Melian Dialogue”)
The summer grasses—
the sole remnants of many
brave warriors’ dreams.Matsuo Bashō
Someone is imprisoned in a room if the door is unlocked, opens inwards; but it doesn’t occur to him to pull, rather than push against it.
Wittgenstein
Sometimes driven aground by the photon storms, by the swirling of the galaxies, clockwise and counterclockwise, ticking with light down the dark sea-corridors lined with our silver sails, our demon-haunted sails, our hundred-league masts as fine as threads, as fine as silver needles sewing the threads of starlight, embroidering the stars on black velvet, wet with the winds of Time that go racing by. The bone in her teeth! The spume, the flying spume of Time, cast up on these beaches where old sailors can no longer keep their bones from the restless, the unwearied universe. Where has she gone? My lady, the mate of my soul? Gone across the running tides of Aquarius, of Pisces, of Aries. Gone. Gone in her little boat, her nipples pressed against the black velvet lid, gone, sailing away forever from the star-washed shores, the dry shoals of the habitable worlds. She is her own ship, she is the figurehead of her own ship, and the captain. Bosun, Bosun, put out the launch! Sailmaker, make a sail! She has left us behind. We have left her behind. She is in the past we never knew and the future we will not see. Put out more sail, Captain, for the universe is leaving us behind...
Hethor (Citadel of the Autarch, Gene Wolfe)
Special knowledge can be a terrible disadvantage if it leads you too far along a path you cannot explain anymore.
Brian Herbert (Dune: House Harkonnen)
Stop hoping you will change the will of the gods by praying.
Virgil (Aeneid)
Streaming in the wind
The smoke from Fuji
Vanishes in the sky;
I know not where
These thoughts of mine go, either.Saigyō
Tell it in the capital:
That like the steadfast pine trees
On Takasago’s sands,
At Onoe, the cherries on the hills
Yet wait in the fullness of their bloom.Fujiwara no Teika
That the pleasure arising to man
from contact with sensible objects,
is to be relinquished because accompanied by pain—
such is the reasoning of fools.
The kernels of the paddy, rich with finest white grains,
What man, seeking his own true interest,
would fling them away
because of a covering of husk and dust?Carvaka
Where death is, I am not; where I am, death is not.
Lucretius

Where is the end of them, the fishermen sailing
Into the wind’s tail, where the fog cowers?
We cannot think of a time that is oceanless
Or of an ocean not littered with wastage
Or of a future that is not liable
Like the past, to have no destination.We have to think of them as forever bailing,
Setting and hauling, while the North East lowers
Over shallow banks unchanging and erosionless
Or drawing their money, drying sails at dockage;
Not as making a trip that will be unpayable
For a haul that will not bear examination.T. S. Eliot (Four Quartets)
I sat upon the shore
Fishing, with the arid plain behind me
Shall I at least set my lands in order?
London Bridge is falling down falling down falling down
Poi s’ascose nel foco che gli affina
Quando fiam uti chelidon—O swallow swallow
Le Prince d’Aquitaine à la tour abolie
These fragments I have shored against my ruins
Why then Ile fit you. Hieronymo’s mad againe.
Datta. Dayadhvam. Damyata.Shantih shantih shantih

Smooth between sea and land
Is laid the yellow sand,And here through summer days
The seed of Adam plays....Here on the level sand,
Between the sea and land,What shall I build or write
Against the fall of night?Tell me of runes to grave
That hold the bursting wave,Or bastions to design
For longer date than mine.Shall it be Troy or Rome
I fence against the foam,Or my own name, to stay
When I depart for aye?Nothing: too near at hand,
Planing the figured sand,Effacing clean and fast
Cities not built to lastAnd charms devised in vain,
Pours the confounding main.A. E. Housman (More Poems, XLV)

Fruits fail and love dies and time ranges;
Thou art fed with perpetual breath,
And alive after infinite changes,
And fresh from the kisses of death.“Dolores (Notre-Dame des Sept Douleurs)”, Algernon Charles Swinburne
For the crown of our life as it closes
Is darkness, the fruit thereof dust;
No thorns go as deep as a rose’s,
And love is more cruel than lust.“Dolores (Notre-Dame des Sept Douleurs)”, Algernon Charles Swinburne
We shall change as the things that we cherish,
Shall fade as they faded before,
As foam upon water shall perish,
As sand upon shore.“Dolores (Notre-Dame des Sept Douleurs)”, Algernon Charles Swinburne
Out of memory.
We wish to hold the whole sky,
But we never will.

Where is the song before it is sung? Where is the dance before it is danced?
Alexander Herzen
Frozen pond reflects
the moon in perfect stillness—
but I toss a stone,
shattering this illusion;
truth is never motionless.GPT-4.5
If at the end of our journey
There is no final
Resting place
Then we need not fear
Losing our way.Ikkyu

Whereof one cannot speak, thereof one must be silent.
All that is gold does not glitter; not all those who wander are lost... but you, evidently, are.

…And you know what’s going to happen now. You should admit your situation.
There would be more dignity in it.
If the rule you followed brought you to this, of what use was the rule?

People can only save themselves.
One person saving another is impossible.

It is universally admitted that the unicorn is a supernatural being and one of good omen; thus it is declared in the Odes, in the Annals, in the biographies of illustrious men, and in other texts of unquestioned authority. Even the women and children of the common people know that the unicorn is a favorable portent. But this animal does not figure among the domestic animals, it is not easy to find, it does not lend itself to any classification. It is not like the horse or the bull, the wolf or the deer. Under such conditions, we could be in the presence of a unicorn and not know with certainty that it is one. We know that a given animal with a mane is a horse, and that one with horns is a bull. We do not know what a unicorn is like.
Jorge Luis Borges, “Kafka And His Precursors” (1951)

One of my insistent pleas to God and my guardian angel was that I not dream of mirrors; I recall clearly that I would keep one eye on them uneasily. I feared sometimes that they would begin to veer off from reality; other times, that I would see my face in them disfigured by strange misfortunes. I have learned that this horror is monstrously abroad in the world again...What dreadful bondage, the bondage of my face—or one of my former faces. Its odious fate makes me odious as well, but I don’t care anymore.
Jorge Luis Borges, “Covered Mirrors” (1934-09-15)

Though both trees and blooms
Have lost their hue,
For the ocean’s
Flowering waves
No autumn comes.Fun’ya no Yasuhide

Death, tho I see him not, is near
And grudges me my eightieth year.
Now, I would give him all these last
For one that fifty have run past.
Ah! he strikes all things, all alike,
But bargains: those he will not strike.“CLXVI. Age”, William Savage Landor
(pg400, The Last Fruit Off an Old Tree, 1853)



Loveliest of trees, the cherry now
Is hung with bloom along the bough,
And stands about the woodland ride
Wearing white for Eastertide.
Now, of my threescore years and ten,
Twenty will not come again,
And take from seventy springs a score,
It only leaves me fifty more.
And since to look at things in bloom
Fifty springs are little room,
About the woodlands I will go
To see the cherry hung with snow.A. E. Housman

III.
Those masterful images because complete
Grew in pure mind but out of what began?
A mound of refuse or the sweepings of a street,
Old kettles, old bottles, and a broken can,
Old iron, old bones, old rags, that raving slut
Who keeps the till. Now that my ladder's gone
I must lie down where all the ladders start
In the foul rag and bone shop of the heart.“The Circus Animals’ Desertion”, William Butler Yeats (Last Poems, 1939)











To my good friend
Would I show, I thought,
The plum blossoms,
Now lost to sight
Amid the falling snow.Yamabe no Akahito (Man’yōshū)

Ignoring its voice,
how many generations of men
have grown old?
Always the same temple’s bells
in the same capital’s mountains.Shōtetsu (“Bells at an Old Temple”)



Along the pathway
the wind of evening
raises its voice.
In the market—no one
but the dust, piling up.Shōtetsu

The gloom of dusk.
An ox from out in the fields
comes walking my way;
and along the hazy road
I encounter no one.Shōtetsu (“An Animal in Spring”)




What are they burning, what are they burning,
Heaping and burning in a thunder-gloom?Now is the time for the burning of the leaves.
They go to the fire; the nostril pricks with smoke
Wandering slowly into a weeping mist.
Brittle and blotched, ragged and rotten sheaves!
A flame seizes the smouldering ruin and bites
On stubborn stalks that crackle as they resist.The last hollyhock’s fallen tower is dust;
All the spices of June are a bitter reek,
All the extravagant riches spent and mean.
All burns! The reddest rose is a ghost;
Sparks whirl up, to expire in the mist: the wild
Fingers of fire are making corruption clean.Now is the time for stripping the spirit bare,
Time for the burning of days ended and done,
Idle solace of things that have gone before:
Rootless hope and fruitless desire are there;
Let them go to the fire, with never a look behind.
The world that was ours is a world that is ours no more.They will come again, the leaf and the flower, to arise
From squalor of rottenness into the old splendour,
And magical scents to a wondering memory bring;
The same glory, to shine upon different eyes.
Earth cares for her own ruins, naught for ours.
Nothing is certain, only the certain spring.“The Burning of the Leaves” § I (1944), Robert Laurence Binyon


Hill upon hill, the road seems to be lost,
Willows and flowers, another village appearsQiu Xiaolong, Death of a Red Heroine
A struck gong resounds—
crows launch from winter branches;
the cold moon behind.
Bronze bell’s lonely voice—
even the crows must depart.
Bright moon through the pines.
Gong strikes the old pond—
crows ripple on night’s mirror.
And the deep, still moon.
Bell’s mouth opens: crows.
Bell’s mouth closes: emptiness.
Moon speaks through the void.
Bell transmutes the night:
silence becomes crows becomes
absence holding light.
The bronze bell speaks once—
making silence into crows
and rushing dark wings;
what is left when they are gone?
Moonlight on winter branches.
At the evening bell,
the crows must flee their old roosts—
boughs left for the wind.
How long our sorrows linger…
the moon remains, still unchanged.
Gong, crows, and motion—
all are waves on time’s surface.
The moon never moves.
Surface: night, bell, wings.
Deeper: silence surrounds all.
Deepest: then the moon.
Each crow carries off
a shard of time’s broken ice—
Moon needs no mirror.
One strike disperses
what seemed eternal—crows flee.
Moon remains, fading.









Webster was much possessed by death
And saw the skull beneath the skin;
And breastless creatures under ground
Leaned backward with a lipless grin.Daffodil bulbs instead of balls
Stared from the sockets of the eyes!
He knew that thought clings round dead limbs
Tightening its lusts and luxuries.“Whispers of Immortality”, T. S. Eliot 1920

“It’s this planet”, Scytale said. “It raises questions.”
“Planets don’t speak!”
“This one does.”
“Oh?”
“It speaks of creation. Sand blowing in the night, that is creation.”
“Sand blowing…”
“When you awaken, the first light shows you the new world—all fresh and ready for your tracks.”
Untracked sand? Edric thought. Creation? He felt knotted with sudden anxiety. The confinement of his tank, the surrounding room, everything closed in upon him, constricted him… “So?”
“Another night comes”, Scytale said. “Winds blow.”
Frank Herbert, Dune Messiah










Smoke spoke in loops.
“I stop when I stop”, the fire said to the last pine.
The pine crackled: “We grow when we grow.”
Ash fell between them. Pause.
GPT-5
The world doesn't end at San Francisco. But some mornings it practices.
GPT-5







“Where has the time gone?”
I wonder in my study,
rain on the windows;
and the years drip down on me
as yellowed papers

Keener sorrow than
the cherry blossoms of spring
is today’s snowfall—
weeping to the ground, leaving
not even a flake behind.

With the melting frost,
this winter my dog departs.
Only snow returns
again and again; and we
too must vanish like the dew.
A stroke of lightning—
none are guaranteed a spring.
Like mountain fires
are the griefs of others:
beautiful, from a distance.(1937–2022)

Other Options
visit the main index
browse documents by tag
browse short blog posts
use a search engine (tips):
Some pages you may have been trying to find: